2. Exact Heston Model Simulations for Characteristic Function Methods#
2.1. Overview#
Exact simulation from Broadie and Kaya [2006]. All implementations in utils.py.
The critical step is inverting the CDF of integrated variance \(\int_0^T V_s \, ds\). We compare COS, Carr-Madan, and CONV for that inversion.
=== Testing COS Method ===
Using cos method for CDF recovery and inversion
Computation time: 0.219 seconds
Min integrated variance: 0.00000000
Values <= 1e-10: 4/204
All positive: False
=== Testing CARR_MADAN Method ===
Using carr_madan method for CDF recovery and inversion
Computation time: 3.942 seconds
Min integrated variance: 0.00000000
Values <= 1e-10: 4/204
All positive: False
=== Testing CONV Method ===
Using conv method for CDF recovery and inversion
Computation time: 9.379 seconds
Min integrated variance: 0.00000000
Values <= 1e-10: 4/204
All positive: False
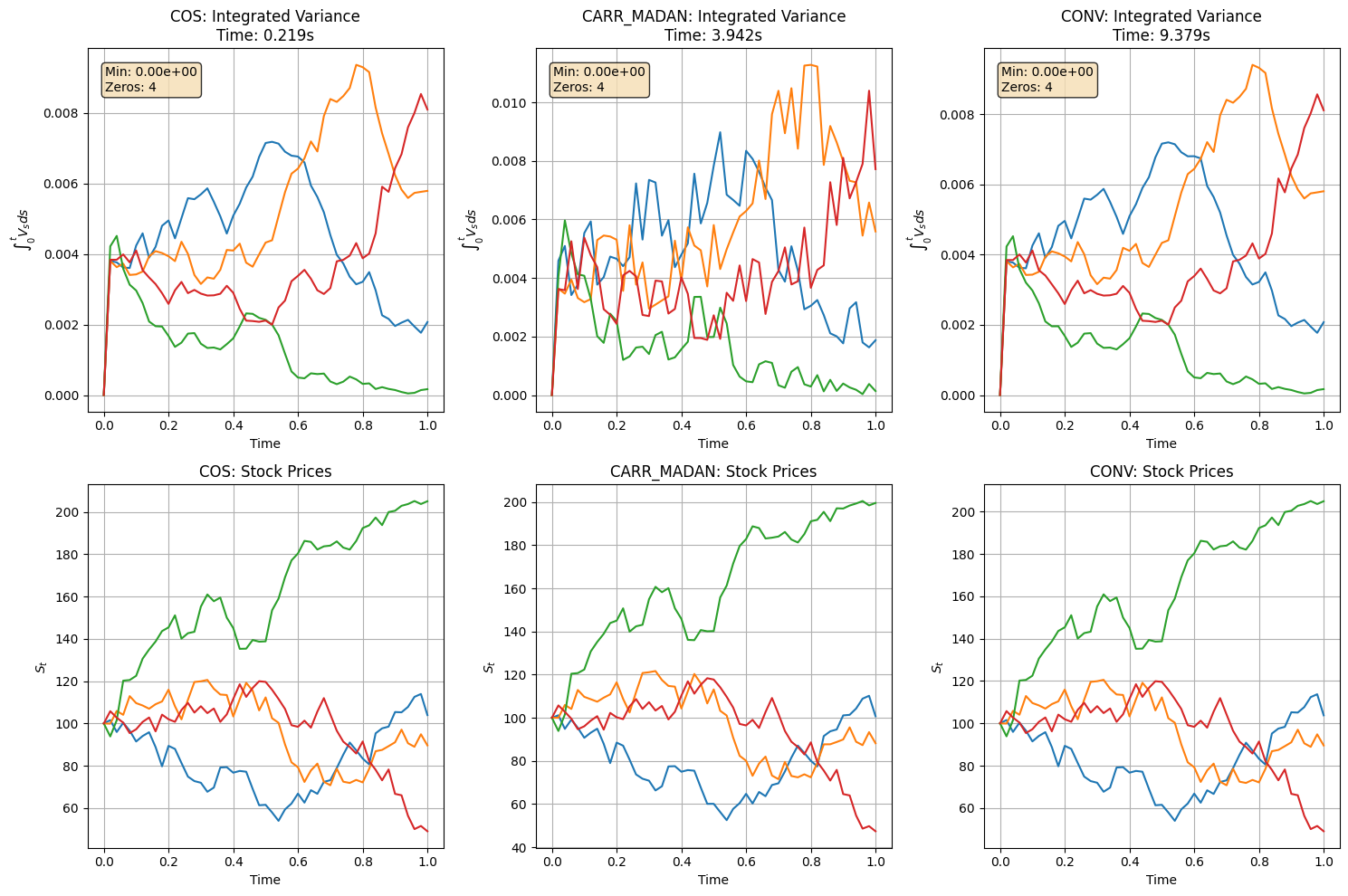
=== Performance Comparison ===
COS: 0.219 seconds
CARR_MADAN: 3.942 seconds
CONV: 9.379 seconds
=== Zero Analysis ===
COS: min=0.00000000, zeros(<=1e-10)=4, exact_zeros=4
CARR_MADAN: min=0.00000000, zeros(<=1e-10)=4, exact_zeros=4
CONV: min=0.00000000, zeros(<=1e-10)=4, exact_zeros=4
Using carr_madan method for CDF recovery and inversion
Min integrated variance: 0.00000000
Zero values: 3/33
All values positive: False
First few values: [[0. 0.00820748 0.01244008 0.0057766 0.00594041]
[0. 0.01216603 0.00488742 0.00421274 0.00721754]
[0. 0.01324573 0.00684682 0.00610527 0.01005191]]
ERROR: Still seeing zeros! The fixed utils.py is not being used.
Please restart the Jupyter kernel and run all cells again.
COS is fastest since it avoids numerical integration. Carr-Madan occasionally produces near-zero PDFs that break Newton’s method — need to investigate. All three produce similar stock trajectories, the difference is in the variance sampling step.
2.2. Comparison#
COS for this application — fast, stable, no integration needed.
2.2.1. Newton-Raphson method vs. brute force approach for CDF inversion#
/home/runner/work/Quantitative-Finance-Book/Quantitative-Finance-Book/char_functions/../utils.py:502: RuntimeWarning: invalid value encountered in scalar multiply
chf = temp1 * temp2 * temp3 / temp4
/home/runner/work/Quantitative-Finance-Book/Quantitative-Finance-Book/char_functions/../utils.py:502: RuntimeWarning: invalid value encountered in scalar divide
chf = temp1 * temp2 * temp3 / temp4
/home/runner/work/Quantitative-Finance-Book/Quantitative-Finance-Book/char_functions/../utils.py:468: RuntimeWarning: invalid value encountered in divide
R
/home/runner/work/Quantitative-Finance-Book/Quantitative-Finance-Book/char_functions/../utils.py:479: RuntimeWarning: invalid value encountered in divide
- R * (1 + np.exp(-R * tau)) / (1 - np.exp(-R * tau))
/home/runner/work/Quantitative-Finance-Book/Quantitative-Finance-Book/char_functions/../utils.py:486: RuntimeWarning: invalid value encountered in divide
np.sqrt(vt * vu)
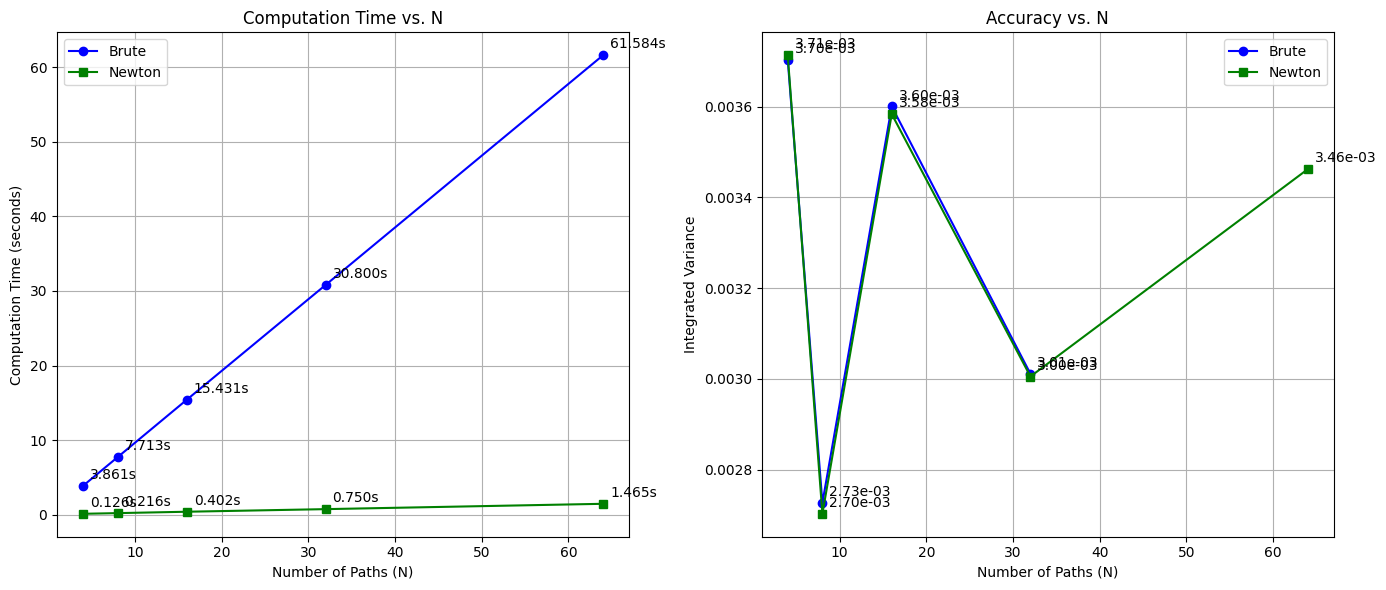
| N | Brute Time (s) | Newton Time (s) | vint_brute_values | vint_newton_values | |
|---|---|---|---|---|---|
| 0 | 4 | 3.860675 | 0.125718 | 0.003704 | 0.003713 |
| 1 | 8 | 7.712639 | 0.215924 | 0.002726 | 0.002702 |
| 2 | 16 | 15.431197 | 0.402222 | 0.003600 | 0.003583 |
| 3 | 32 | 30.799817 | 0.750099 | 0.003011 | 0.003005 |
| 4 | 64 | 61.583629 | 1.465462 | NaN | 0.003462 |