1. Recovering PDF/CDF from Characteristic Functions#
If the characteristic function is known, we can recover the PDF and CDF without needing the density in closed form. The characteristic function of a random variable \(X\):
is the Fourier transform of the density. Inverting it gives us back the PDF:
This is useful for option pricing when we have an analytical CF (Black-Scholes, Heston, Variance-Gamma, etc.) but no closed-form density. We can price options by computing:
Three methods for this inversion: COS, Carr-Madan, and CONV. Each has different convergence properties. COS converges exponentially (N~64 for machine precision on smooth densities), Carr-Madan and CONV algebraically (N~256+).
1.1. COS Method#
The COS method Fang and Oosterlee [2008] expands the density as a Fourier cosine series on \([a,b]\):
The coefficients \(A_k\) can be computed directly from the characteristic function:
No numerical integration needed — just evaluate the CF at specific frequencies. For smooth densities, convergence is exponential: \(O(\rho^N)\) with \(\rho < 1\). With N=64 you often hit machine precision.
The CDF follows by integrating the series:
where \(\omega_k = k\pi/(b-a)\).
Domain \([a,b]\) should capture most of the probability mass — typically \([\mu - L\sigma, \mu + L\sigma]\) with \(L \approx 10\).
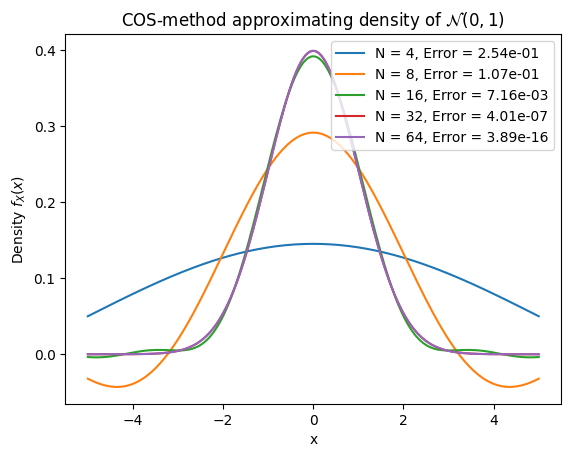
Standard normal density recovery — sweep N from 4 to 64 on \([-10, 10]\) and measure max absolute error.
Convergence table:
N Error CPU time (msec.) Diff. in CPU (msec.)
0 4 2.536344e-01 0.073695 NaN
1 8 1.074376e-01 0.034595 -0.039101
2 16 7.158203e-03 0.052738 0.018144
3 32 4.008963e-07 0.101948 0.049210
4 64 3.885781e-16 0.181580 0.079632
Testing CDF recovery:
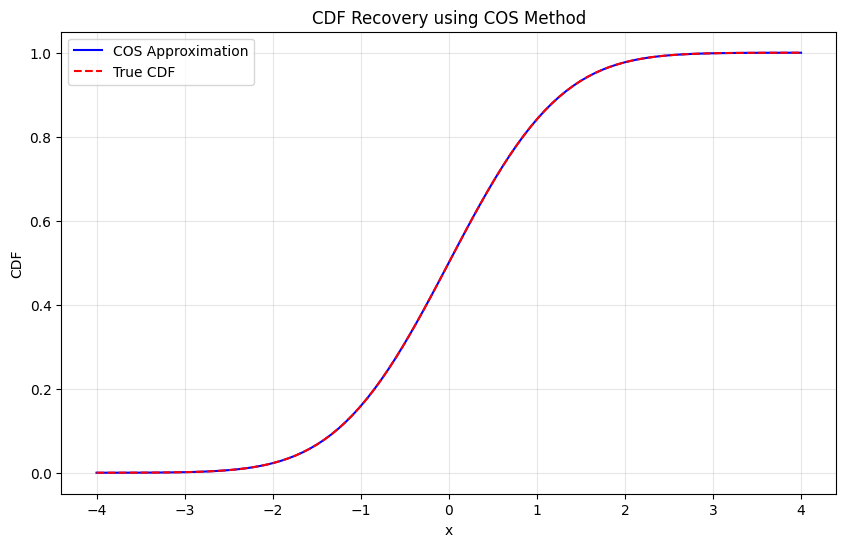
Maximum CDF error: 4.44e-16
1.2. Carr-Madan Method#
Uses Gil-Pelaez inversion. The CDF:
The PDF via inverse Fourier transform:
For numerical stability (especially heavy tails), add a damping factor \(\alpha > 0\):
Then recover: \(f(x) = e^{\alpha x} f_{\alpha}(x)\).
Truncate to \([-u_{\max}, u_{\max}]\) and use FFT for \(O(N \log N)\) complexity. Convergence is algebraic: \(O(N^{-p})\) where \(p\) depends on how fast \(|\varphi(u)|\) decays.
Damping \(\alpha \in [0.5, 1.5]\) works for most cases. Sensitive to parameter choice — need to tune \(\alpha\) and \(u_{\max}\).
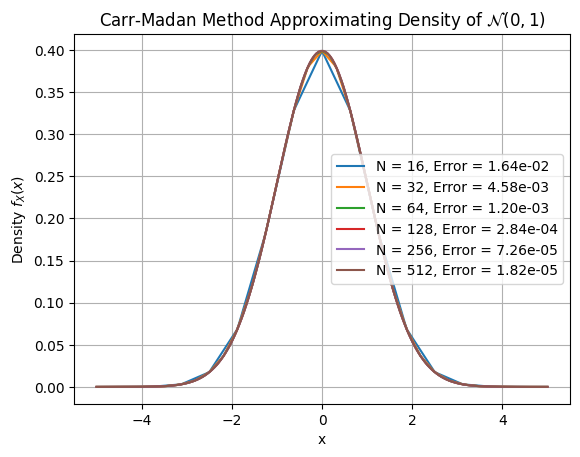
Same test as COS. Carr-Madan needs more grid points since convergence is algebraic.
Convergence plot and table:
N Error CPU time (msec.) Diff. in CPU (msec.)
0 16 0.016351 0.107002 NaN
1 32 0.004577 0.079298 -0.027704
2 64 0.001203 0.074434 -0.004864
3 128 0.000284 0.107098 0.032663
4 256 0.000073 0.051022 -0.056076
5 512 0.000018 0.072336 0.021315
Testing CDF recovery:
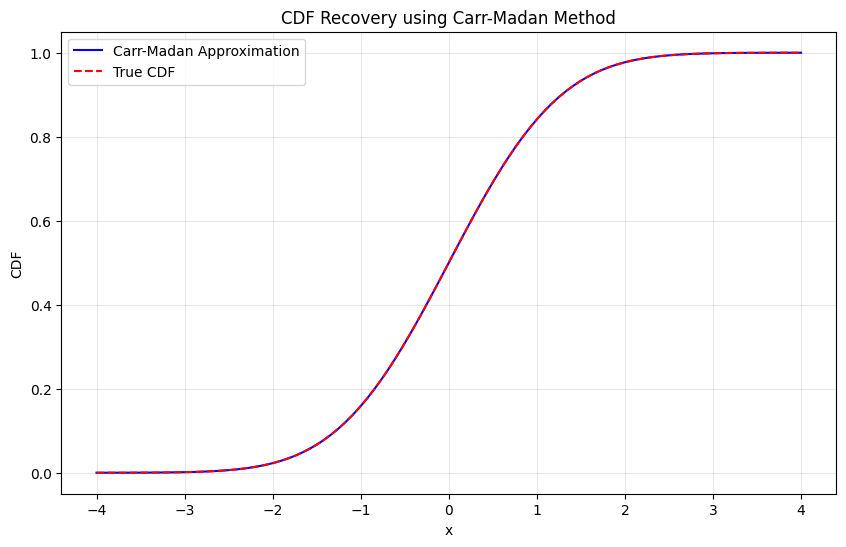
Maximum CDF error: 1.27e-10
1.3. CONV Method#
Discretizes the inverse Fourier transform and uses FFT. Same underlying formula as Carr-Madan:
Grid setup:
Spatial: \(x_k = x_{\min} + k \cdot \Delta x\)
Frequency: \(u_j = j \cdot \Delta u\) where \(\Delta u = 2\pi / L\) and \(L = x_{\max} - x_{\min}\)
The discrete approximation:
This is just an inverse DFT — use np.fft.fft with proper scaling.
Add damping \(\alpha\) for stability with heavy tails, same as Carr-Madan. The phase correction \(e^{-iu_j x_{\min}}\) shifts the output to the desired spatial domain.
Complexity: \(O(N \log N)\). More robust than Carr-Madan for unknown distributions. Convergence is spectral for smooth CFs, algebraic otherwise.
Same test. CONV uses FFT with phase correction — similar to Carr-Madan but different grid handling.
Testing PDF recovery:
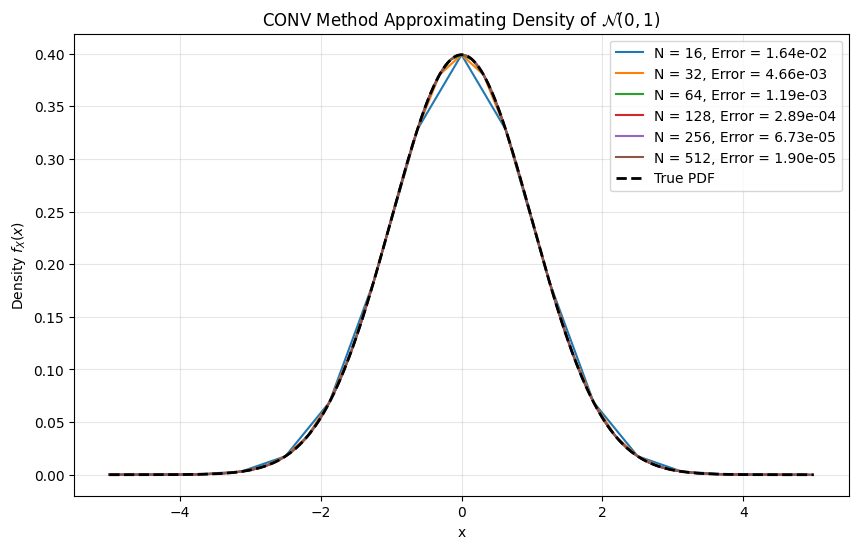
Convergence table:
N Error CPU time (msec.) Diff. in CPU (msec.)
0 16 0.016413 0.118780 NaN
1 32 0.004663 0.104642 -0.014138
2 64 0.001192 0.104499 -0.000143
3 128 0.000289 0.109625 0.005126
4 256 0.000067 0.112200 0.002575
5 512 0.000019 0.184894 0.072694
Testing CDF recovery:
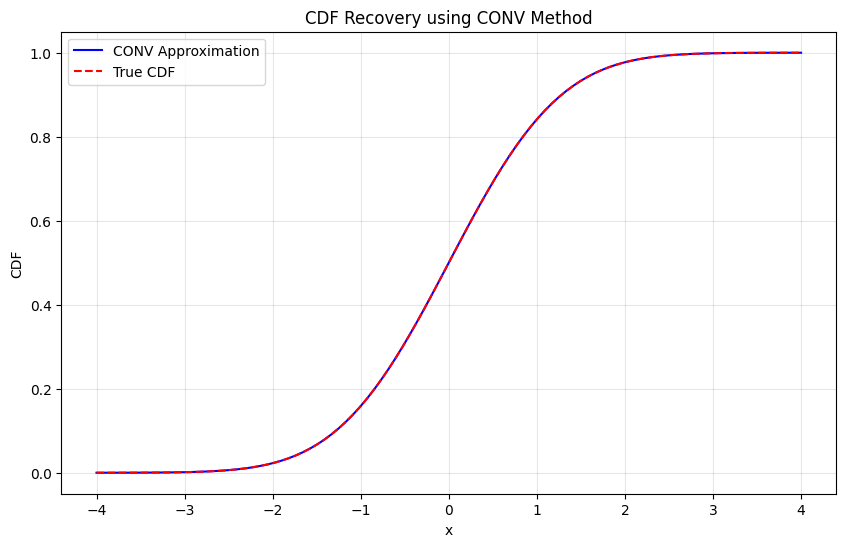
Maximum CDF error: 7.56e-07
======================================================================
CONVERGENCE COMPARISON: Maximum Absolute Error
======================================================================
Method CONV COS Carr-Madan
N
16 0.016413 7.172372e-03 0.016416
32 0.004663 4.032330e-07 0.004663
64 0.001192 3.608225e-16 0.001193
128 0.000289 3.608225e-16 0.000289
256 0.000067 3.608225e-16 0.000067
======================================================================
PERFORMANCE COMPARISON: CPU Time (milliseconds)
======================================================================
Method CONV COS Carr-Madan
N
16 0.102997 0.162315 0.036621
32 0.098491 0.212002 0.033498
64 0.100017 0.398946 0.035882
128 0.107312 0.782943 0.039887
256 0.120211 1.553893 0.046682
| N | Method | Error | CPU time (ms) | |
|---|---|---|---|---|
| 0 | 16 | COS | 7.172372e-03 | 0.162315 |
| 1 | 16 | Carr-Madan | 1.641613e-02 | 0.036621 |
| 2 | 16 | CONV | 1.641260e-02 | 0.102997 |
| 3 | 32 | COS | 4.032330e-07 | 0.212002 |
| 4 | 32 | Carr-Madan | 4.663137e-03 | 0.033498 |
| 5 | 32 | CONV | 4.662527e-03 | 0.098491 |
| 6 | 64 | COS | 3.608225e-16 | 0.398946 |
| 7 | 64 | Carr-Madan | 1.192675e-03 | 0.035882 |
| 8 | 64 | CONV | 1.192463e-03 | 0.100017 |
| 9 | 128 | COS | 3.608225e-16 | 0.782943 |
| 10 | 128 | Carr-Madan | 2.887765e-04 | 0.039887 |
| 11 | 128 | CONV | 2.886811e-04 | 0.107312 |
| 12 | 256 | COS | 3.608225e-16 | 1.553893 |
| 13 | 256 | Carr-Madan | 6.734658e-05 | 0.046682 |
| 14 | 256 | CONV | 6.730085e-05 | 0.120211 |
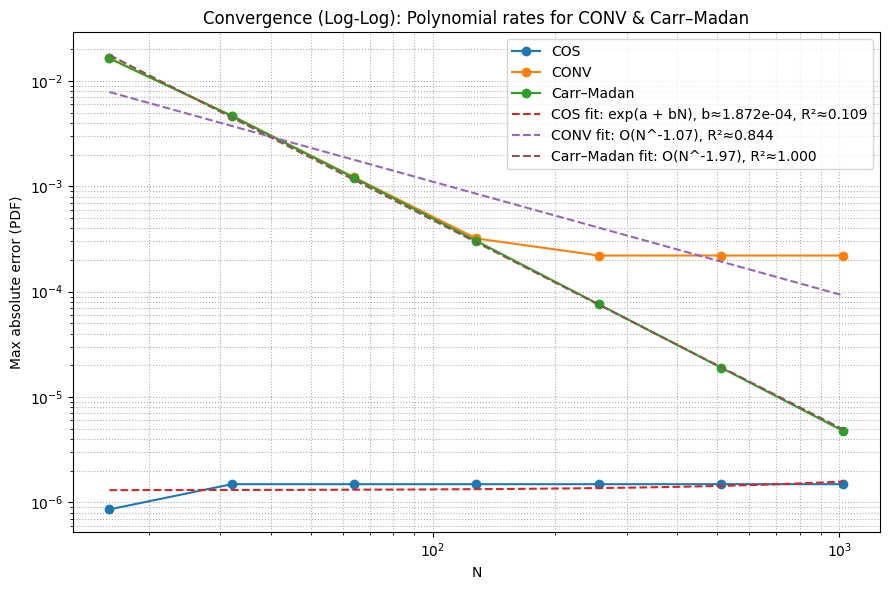
=== Empirical Convergence Summary ===
COS: last N=1024, error=1.487e-06, R²≈0.109
CONV: last N=1024, error=2.206e-04, R²≈0.844
Carr–Madan: last N=1024, error=4.753e-06, R²≈1.000
From the convergence plots: COS reaches machine precision at N~64 for the standard normal, while Carr-Madan and CONV need N~256+. COS is the obvious choice for smooth distributions. Carr-Madan handles heavier tails better. CONV is the most robust but slowest.
TODO: compare on heavier-tailed distributions (Variance Gamma, NIG) where the COS advantage may shrink.